inCNV: An Integrated Analysis Tool for Copy Number Variation on Whole Exome Sequencing - Saowwapark Chanwigoon, Sakkayaphab Piwluang, Duangdao Wichadakul, 2020
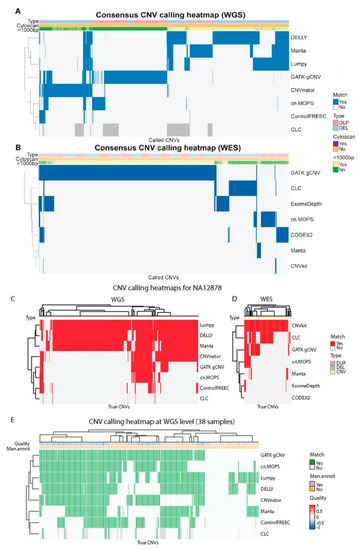
Cancers | Free Full-Text | A Comparison of Tools for Copy-Number Variation Detection in Germline Whole Exome and Whole Genome Sequencing Data

inCNV: An Integrated Analysis Tool for Copy Number Variation on Whole Exome Sequencing | Semantic Scholar
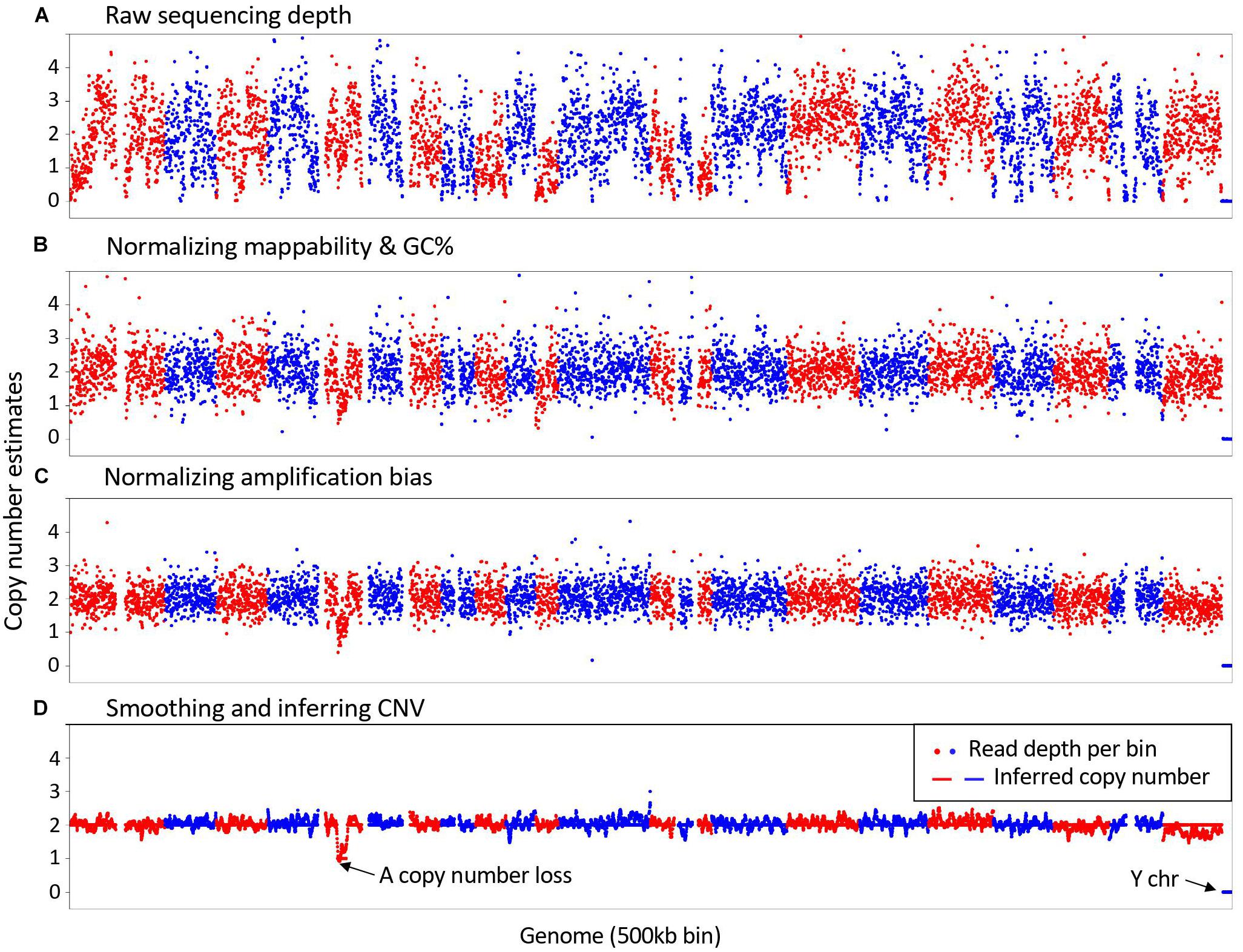
Frontiers | SCCNV: A Software Tool for Identifying Copy Number Variation From Single-Cell Whole-Genome Sequencing
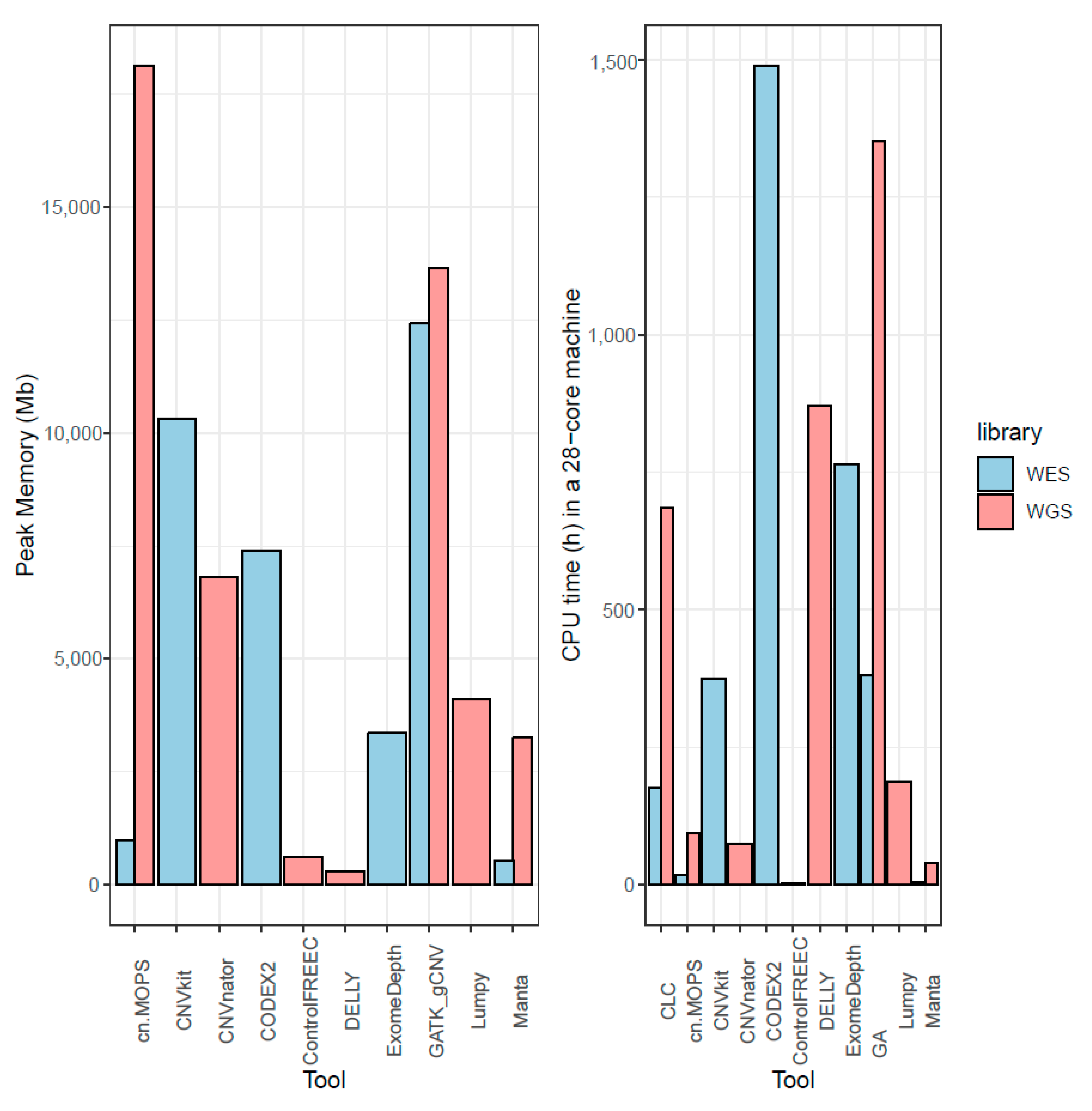
Cancers | Free Full-Text | A Comparison of Tools for Copy-Number Variation Detection in Germline Whole Exome and Whole Genome Sequencing Data
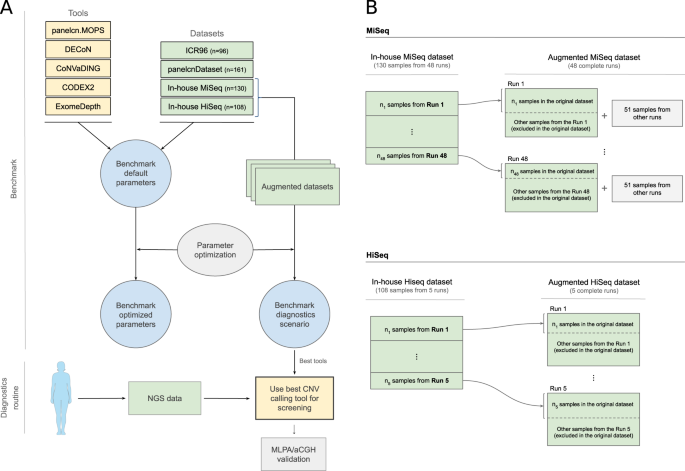
Evaluation of CNV detection tools for NGS panel data in genetic diagnostics | European Journal of Human Genetics
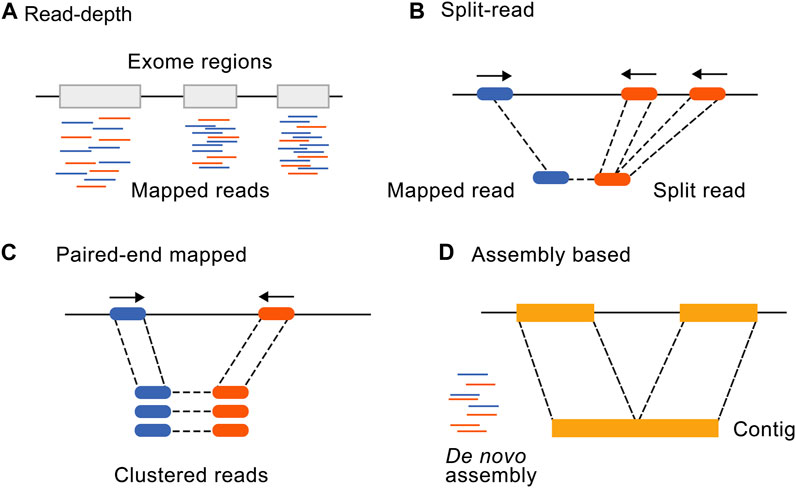
Frontiers | Incorporating CNV analysis improves the yield of exome sequencing for rare monogenic disorders—an important consideration for resource-constrained settings

Comparative study of whole exome sequencing-based copy number variation detection tools | BMC Bioinformatics | Full Text
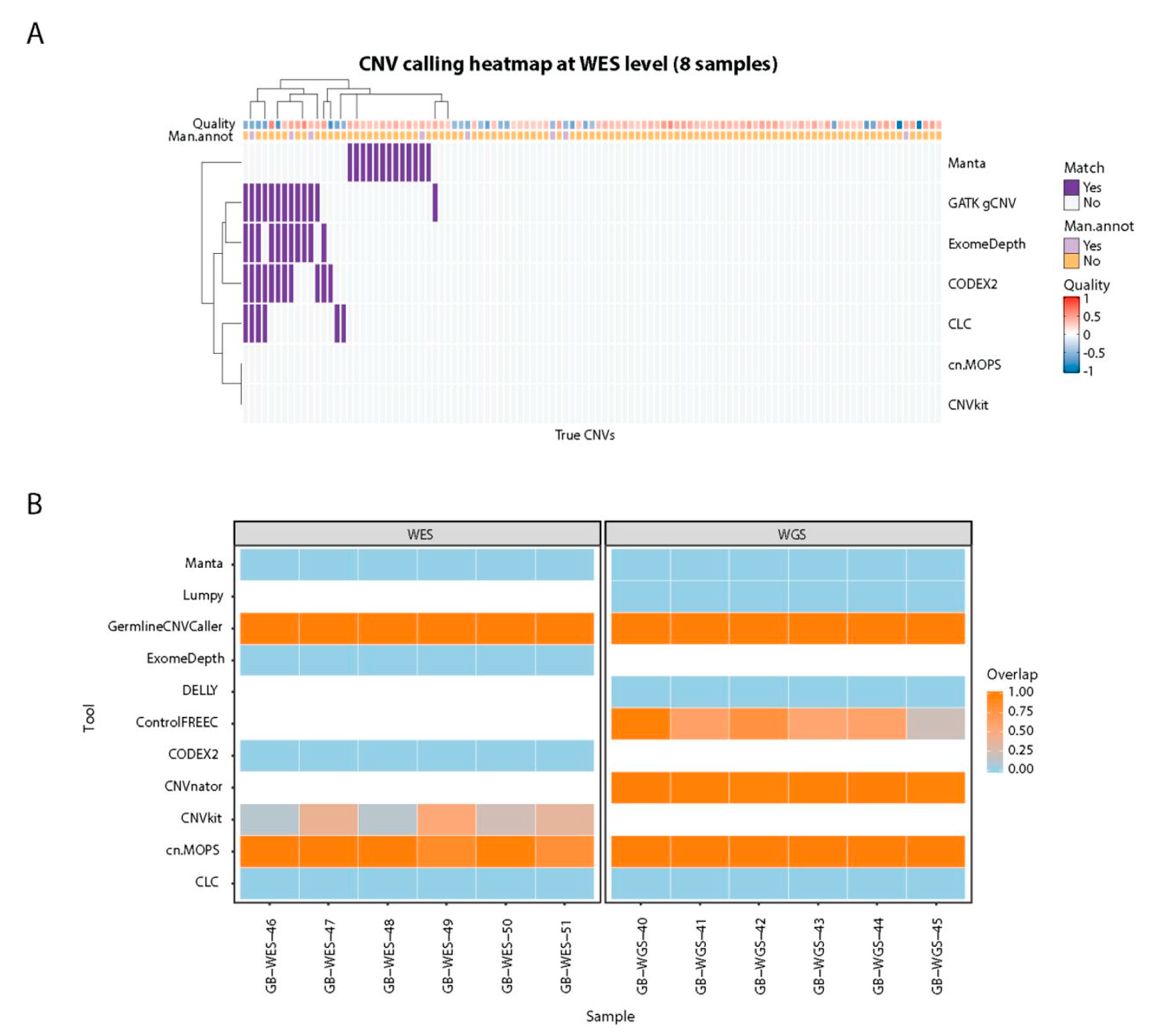
Cancers | Free Full-Text | A Comparison of Tools for Copy-Number Variation Detection in Germline Whole Exome and Whole Genome Sequencing Data

inCNV: An Integrated Analysis Tool for Copy Number Variation on Whole Exome Sequencing - Saowwapark Chanwigoon, Sakkayaphab Piwluang, Duangdao Wichadakul, 2020